- Home
- About
- Partners
- Newcastle University
- University of L'Aquila
- University of Manchester
- Alacris Teranostics GmbH
- University of Pavia
- Polygene
- Consiglio Nazionale delle Ricerche
- INSERM
- Certus Technology
- Charité Universitaet Medizin
- GATC Biotech
- University Medical Center Hamburg Eppendorf
- Evercyte GmbH
- University Hospital of Cologne
- PRIMM Srl
- University of Freiburg
- University of Antwerp
- Finovatis
- Research
- SYBIL at a glance
- Bone
- Growth plate
- Desbuquois dysplasia
- Diastrophic dysplasia
- MCDS
- Osteopetrosis
- Osteoporosis
- Osteogenesis imperfecta
- Prolidase deficiency
- PSACH and MED
- Systems biology
- SOPs
- Alcian Blue staining
- Bone measurements
- BrdU labelling
- Cell counting using ImageJ
- Chondrocyte extraction
- Cre genotyping protocol
- DMMB assay for sulphated proteoglycans
- Densitometry using ImageJ
- Double immunofluorescence
- Electron microscopy of cartilage - sample prep
- Extracting DNA for genotyping
- Grip strength measurement
- Histomorphometry on unon-decalcified bone samples
- Immunocytochemistry
- Immunofluorescence
- Immunohistochemistry
- Quantitative X-ray imaging on bones using Faxitron and ImageJ
- Skeletal preps
- TUNEL assay (Dead End Fluorimetric Kit, Promega)
- Toluidine Blue staining
- Toluidine Blue staining
- Von Kossa Gieson staining
- Wax embedding of cartilage tissue
- Contact Us
- News & Events
- Links
- Portal
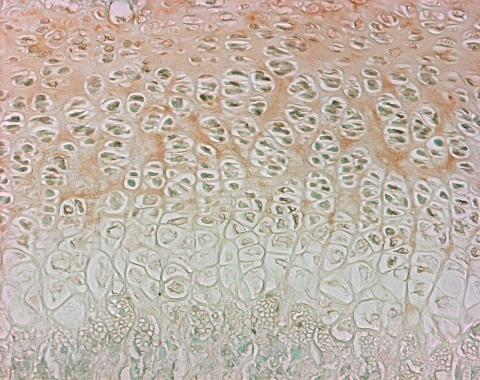
Immunohistochemistry
Sample preparation
- sacrifice the animals and dissect knee samples
- fix in 95%EtOH 5%HAc over 48h in the fridge
- decalcify in 20% EDTApH7.4 for 2 weeks (shaking)
- wax embed and section (6um sections)
Note
to generate reliable data, this experiment should be performed on matched sections from 3 unrelated animals per genotype
Staining steps
dewax in xylene 2 x 5min
↓
100% EtOH 3min
↓
90% EtOH 3min
↓
70% EtOH 3min
↓
50% EtOH 3min
↓
dH2O 2x 3min (from here on, do not let the samples dry!)
↓
1x PBS 2x 3min
↓
225ml MetOH + 7.5ml H2O2 30min (quench endogenous peroxidase)
↓
1x PBS 3x 5min
↓
mark the area around the sections with ImmEdge pen
keep the slides in a darkened humidified chamber during all incubations
↓
0.2% bovine hyaluronidase in 1x PBS (antigen unmasking) 30min or 15 min at 37ºC
↓
1x PBS 3x 5min
↓
block in 10ml PBS/BSA (1% BSA in 1x PBS) + 60ul serum 1h
↓
1ºAb in PBS/BSA 1h
↓
PBS/BSA 2x 5min
↓
2ºAb in PBS/BSA/serum 1h
↓
prepare ABC reagent (2.5ml PBS+dropA+dropB, 30min before use)
↓
1x PBS 3x 5min
↓
ABC reagent 30min
↓
1x PBS 2x 5min
↓
DAB (prepare fresh) 2-10min, stop in dH2O
↓
methyle green 10 min
↓
dunk in 3 fresh changes of tap water
↓
95% EtOH 3min
↓
100% EtOH 5min
↓
dehydrate in xylene 2x 5min
. ↓
coverslip, dry overnight
Modifications
- if using a fluorescent secondary, skip the peroxidase quenching step and mount in Vectashield straight after washing off the excess of the 2º antibody
- other unmasking methods: proteinase K (20 ug/ml in PBS; 37ºC 10min), citrate buffer boil (10mM citric acid, pH 6.0, 0.05% Tween; microwave on high power 3-4min, medium power 3-4 min, low power 3-4min; allow to cool down on bench)
end result: positive staining brown, nuclei green